Introduction
This project is an amalgamation of extensive research and practical laboratories focusing on the mathematical foundations of Artificial Intelligence. It bridges the gap between theoretical math (Linear Algebra, Calculus, Probability) and practical implementation in Machine Learning and Deep Learning.
The research spans from fundamental dimensionality reduction techniques to state-of-the-art multi-modal Vision-Language Models.
Part 1: PCA and Clustering (Lab 2)
We implemented Principal Component Analysis (PCA) from scratch to understand the mathematics behind dimensionality reduction before applying clustering algorithms.
Mathematical Foundation of PCA
PCA transforms data from its original space to a new space with uncorrelated principal components, retaining the maximum variance.
- Z-score Standardization: We first normalize the data:
- Covariance Matrix: We compute the covariance matrix to understand the relationship between variables:
- Eigen Decomposition: We find the eigenvalues and eigenvectors of where . The eigenvectors corresponding to the largest eigenvalues represent the principal components.
Note (Evaluation Metrics)
We used Explained Variance Ratio (EVR) and Cumulative Explained Variance Ratio (CEVR) to determine the optimal number of principal components to retain (usually aiming for CEVR).
Clustering Applications
After applying PCA, we clustered the reduced data using two unsupervised learning algorithms:
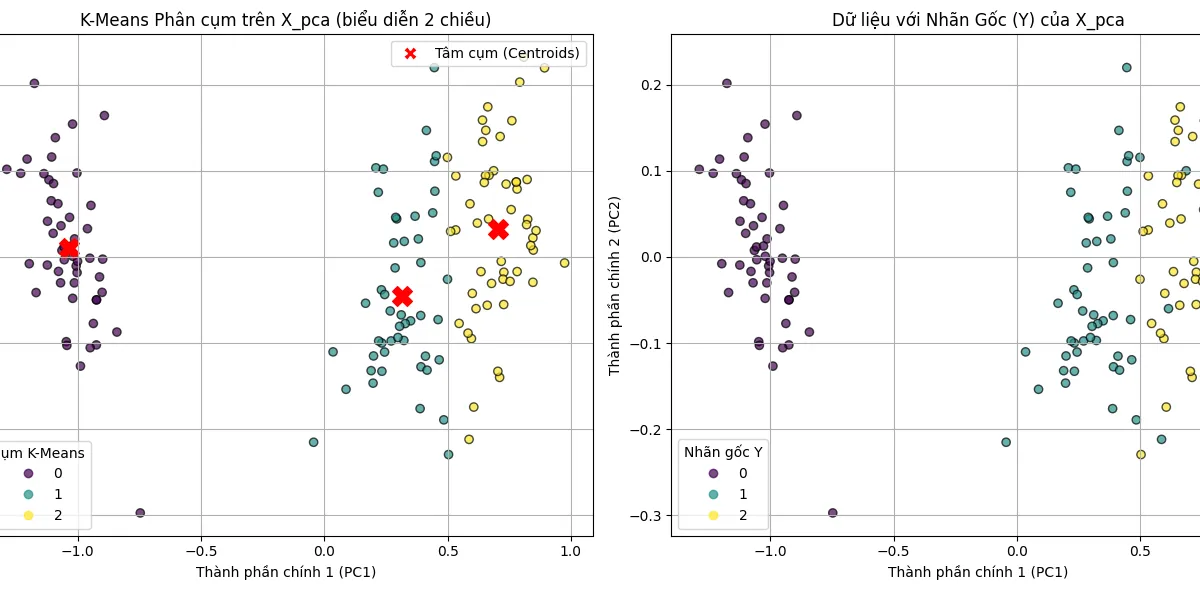
- K-Means Clustering: Assigns data points to clusters by minimizing the sum of squared distances to the cluster centroids.
- Gaussian Mixture Model (GMM): Represents data as a mixture of multivariate Gaussian distributions, trained using the Expectation-Maximization (EM) algorithm to find the Maximum Likelihood Estimate (MLE).
Warning (Results on ABIDE II Dataset)
When applied to the complex ABIDE II dataset (autism brain imaging), K-Means achieved accuracy and GMM achieved . The low F1-scores highlight the limitations of purely mathematical/statistical transformations on highly complex medical data without deep learning architectures like Graph Convolutional Networks (GCN).
Part 2: Contrastive Language-Image Pretraining (CLIP)
The final project shifted focus to advanced Deep Learning by deeply researching OpenAI’s CLIP model, a foundation model that connects computer vision and natural language processing.
The Power of Natural Language Supervision
Unlike traditional models trained on fixed label sets (e.g., ImageNet’s 1000 classes), CLIP is trained on WIT (WebImageText), a dataset of 400 million image-text pairs. It uses natural language supervision, allowing it to learn highly generalized representations and perform zero-shot classification on completely unseen datasets.
Architecture
CLIP uses a dual-encoder architecture:
- Image Encoder: Uses either ResNet (with anti-aliased blur pooling and attention pooling modifications) or Vision Transformer (ViT).
- Text Encoder: Uses a Transformer decoder with Masked Self-Attention and Byte Pair Encoding (BPE).
Contrastive Learning
Instead of predicting the exact text caption (generative), CLIP uses a Symmetric Cross-Entropy Loss to maximize the cosine similarity between the correct image-text pairs in a batch while minimizing it for the incorrect pairs.
Solution (Application: Face Recognition)
We applied a pretrained CLIP (ViT-B/32) as a feature extractor combined with FAISS k-NN for face recognition on the Labeled Faces in the Wild dataset. This approach achieved significantly better accuracy and generalization ( Accuracy) compared to fine-tuning traditional CNNs like ResNet or MobileNet from scratch, proving the robustness of CLIP’s multi-modal embedding space.
Model Comparisons
We also researched and compared CLIP against similar models:
- ALIGN: Scales up using 1.8 billion highly noisy image-text pairs, proving that massive scale can compensate for noisy labels.
- BLIP: Introduces “CapFilt” to bootstrap the dataset (generating synthetic captions and filtering noisy ones) and uses a unified MED architecture to support both understanding (retrieval) and generation (captioning) tasks.